| Research Article | ||
J. Microbiol. Infect. Dis., (2026), Vol. 16(1): 28–36 Research Article Genomic and phenotypic characterization of multidrug-resistant Klebsiella pneumoniae in neonatal care settingsMohamed T. Saad1, Nadia E. Sifennasr2, Mahmoud B. Agena3, Omar S. Elhenshir1, Ahmed A. Zaghdani1, Abdlrhman M. Alsonosi4, Budour R. Elmihub1 and Khaled M. Ibrahim1*1Microbiology Department, Libyan Biotechnology Research Centre, Tripoli, Libya 2Department of Medical Microbiology and immunology, Faculty of Medicine, University of Tripoli, Tripoli, Libya 3Microbiology Department, Libyan Medical Research Centre, Az-Zawia, Libya 4Department of Medical Microbiology, Faculty of Medicine, Sebha University, Sebha, Libya *Corresponding Author: : Khaled Ibrahim. Microbiology Department, Libyan Biotechnology Research Centre, Tripoli, Libya. Email: ntu.telesi [at] gmail.com Submitted: 12/12/2025 Revised: 02/02/2026 Accepted: 11/02/2026 Published: 28/02/2026 © 2026 Open Veterinary Journal
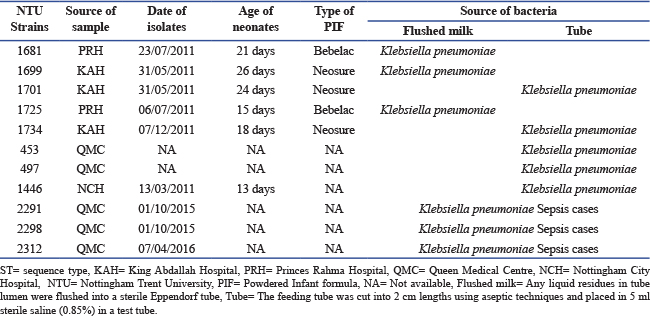
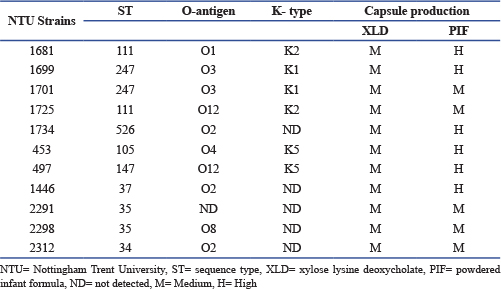
ABSTRACTBackground: Klebsiella pneumoniae, classified by the World Health Organization as a critical priority pathogen, poses a major global health threat. Klebsiella pneumoniae causes a wide range of infections, including sepsis, urinary tract infections, meningitis, soft tissue infections, liver abscesses, and pneumonia. Its pathogenicity is driven by virulence and antimicrobial resistance genes that complicate treatment and infection control. Aim: This study aimed to evaluate the virulence-linked traits of eleven K. pneumoniae isolates associated with neonatal infections and to compare them with two reference genomes (NTUH K2044 and MGH 78578) to assess virulence gene linkage and pathogenic potential. Methods: Eleven neonatal K. pneumoniae isolates (six from sepsis cases and five from feeding tubes) were evaluated. Isolates were characterised using standard microbiological methods, sequence typing, and whole genome sequencing (Illumina MiSeq). Efflux pump genes representing five families were compared between the neonatal isolates and the reference genomes, which was based on the presence/absence of genes only. Results: Whole genome sequencing revealed eight sequence types (ST34, ST35, ST37, ST105, ST111, ST147, ST247, and ST526) and six O-antigen serogroups (O1, O2, O3, O4, O8, and O12). Phenotypically, all represented isolates showed high amount of capsular production on powdered infant formula media than on xylose lysine deoxycholate media. Antibiotic resistance profiles of the presented isolates were resistant to 10 from 11 antibiotics tested except ciprofloxacin, and were confirmed Extended spectrum β-lactamase producers. Comparative analysis showed that the K. pneumoniae isolates carried a set of efflux pump-associated genes that may be linked to multidrug resistance and virulence-related potential, compared with the reference isolates NTUH K2044 and MGH 78578. Conclusion: The findings highlight the genomic diversity and multidrug-resistant nature of neonatal K. pneumoniae isolates and support the use of comparative genomic analyses to improve understanding of resistance and virulence-associated features in neonatal care settings. Keywords: Klebsiella pneumoniae, Efflux pump genes, Sepsis, Feeding tubes and neonates. IntroductionKlebsiella pneumoniae is a Gram-negative opportunistic pathogen and the leading cause of both community-acquired and healthcare-associated infections worldwide (Santajit and Indrawattana 2016; De Oliveira et al., 2020). According to recent reports from the World Health Organization, K. pneumoniae is classified within the “critical priority” group of pathogens, for which antimicrobial resistance represents a significant global threat and the development of new therapeutic strategies is urgently required (n.d ). Members of the genus Klebsiella are responsible for a substantial proportion of hospital-acquired infections, including pneumonia, bloodstream infections, urinary tract infections, surgical site infections, and device-associated infections. These infections disproportionately affect vulnerable individuals, particularly neonates, immunocompromised patients, and the elderly (Navon-Venezia et al., 2017). The polysaccharide capsule of Klebsiella spp. is a key virulence-associated feature that can enhance survival by protecting against environmental stresses and host immune defences. However, the role of the capsule in pathogenicity is context-dependent and varies with isolate background and host conditions, and its presence alone does not necessarily predict virulence (Ogrodzki and Forsythe 2015). The O and K serotypes of K. pneumoniae have substantial clinical and epidemiological importance (Hansen et al., 2002; Hsieh et al., 2012). Notably, the K1 serotype, recently recognized as a hypervirulent variant, is strongly associated with pyogenic liver abscesses and severe metastatic complications, including septic ocular and central nervous system infections. Similarly, the O1 serotype has been linked to hypervirulent isolates that cause pyogenic liver abscesses. In addition, both O and K serotypes play a critical role in defining K. pneumoniae clonality during investigations of nosocomial outbreaks (Choby and Howard-Anderson 2020; Struve et al., 2015). Previous studies have reported that the O1 and O2 serotypes collectively account for more than 65% of human Klebsiella infections, with the O1 serotype being more prevalent than O2 (Fang et al., 2016; Heinz et al., 2016). Among the diverse mechanisms contributing to multidrug resistance in K. pneumoniae, efflux pump systems remain comparatively underexplored yet, although they play a crucial role. These transport systems belong to several major families, including the resistance nodulation division (RND) family, which is often associated with high capacity multidrug export; the multidrug and toxic compound extrusion (MATE) family, typically involved in the removal of drugs and toxic compounds; the major facilitator superfamily (MFS), which comprises numerous single gene or operon based transporters with variable substrate specificity; the small multidrug resistance (SMR) family, which consists of relatively small transporters that contribute to resistance despite limited substrate ranges; and the ATP binding cassette (ABC) family, which includes energy dependent transporters that, although more common in Gram positive bacteria, are also present in Gram negative bacteria. The hypervirulent K. pneumoniae isolate NTUH K2044, originally isolated from a community-acquired pyogenic liver abscess, exemplifies the extensive efflux pump repertoire that can be present in this species, harbouring more than 15 efflux pump-related open reading frames across multiple families, including AcrAB and KexD RND, KmrA MFS, and KpnEF SMR. Several studies have reported that functional analyses have demonstrated that disruption of selected efflux pump genes increases susceptibility to multiple classes of antibiotics, to detergents, and to agents that induce cell envelope stress (Piddock 2006a; Piddock 2006b; Ni et al., 2020; Fursova et al., 2021). In addition to mediating antibiotic resistance, efflux pumps also contribute to tolerance against non-antibiotic compounds, such as antiseptics and disinfectants, thereby facilitating persistence on hospital surfaces and medical devices, an attribute of particular relevance in clinical environments (Ramos et al., 2014; Wyres and Holt 2018). Two fully sequenced K. pneumoniae reference isolates, NTUH K2044 and MGH 78578, are widely used in comparative genomic analyses. NTUH K2044 represents a hypervirulent isolate, community-associated isolate, whereas MGH 78578 is a classical hospital-associated isolate. These genomes provide valuable benchmarks for evaluating virulence determinants, efflux pump gene repertoires, antimicrobial resistance genes, and capsular and adhesive features. Comparative analyses relative to these reference isolates can provide insights into the evolutionary trajectories of classical, hypervirulent, and multidrug-resistant lineages, as well as the genetic factors associated with increased pathogenicity and clinical severity (Davin-Regli et al., 2016; Piddock 2006; Joshi et al., 2025). Aim of the studyThis study aimed to characterise eleven K. pneumoniae isolates, including six recovered from neonatal sepsis cases and five from neonatal feeding tubes. Using phenotypic analyses of capsular production and antimicrobial susceptibility, alongside sequence typing, O-antigen serogroup determination, and whole genome sequencing (Illumina MiSeq), we investigated the distribution of virulence and resistance-associated genes. Particular emphasis was placed on assessing the presence and diversity of efflux pump genes across multiple transporter families. Comparative genomic analyses with the reference isolates NTUH K2044 and MGH 78578 were performed to evaluate relative virulence potential, the convergence of virulence and antimicrobial resistance, and the potential risk posed to neonatal health. Identification of efflux pump-associated virulence factors was based solely on the presence/ absence of the corresponding genes. Materials and MethodsBacterial isolatesThe study was conducted in 2018, “All isolates, together with positive and negative control strains, were preserved in TSB containing 80% glycerol (Thermo Fisher Scientific, UK) at −80°C or −20°C for long-term storage. When required, strains were recovered from frozen stocks, subcultured on Tryptone Soya Agar (TSA), and incubated aerobically at 37°C for 18 hours. For short-term use, cultures were maintained on TSA at 4°C.” A total of eleven K. pneumoniae isolates were included in this study. Six isolates (453, 497, 1,446, 2,291, 2,298, and 2,312) were obtained from neonates diagnosed with sepsis in two hospitals in Nottingham, United Kingdom. The remaining five isolates (1,681, 1,725, 1,701, 1,699, and 1,734) were recovered from enteral feeding tubes (EFTs) in two hospitals in Jordan (Table 1). Table 1. Klebsiella pneumoniae isolates selected for a comparison study.
ST=sequence type, KAH=King Abdallah Hospital, PRH=Princes Rahma Hospital, QMC=Queen Medical Centre, NCH=Nottingham City Hospital, NTU=Nottingham Trent University, PIF=Powdered Infant formula, NA=Not available, Flushed milk=Any liquid residues in tube lumen were flushed into a sterile Eppendorf tube, Tube=The feeding tube was cut into 2 cm lengths using aseptic techniques and placed in 5 ml sterile saline (0.85%) in a test tube. Capsular production assayCapsule production was assessed using two different growth media: xylose lysine deoxycholate (XLD) agar (Thermo Fisher Scientific, UK) and powdered infant formula (PIF) agar. To prepare PIF agar, 800 ml of Plate Count Agar (PCA) (Thermo Fisher Scientific, UK) was prepared according to the manufacturer’s instructions. Subsequently, 200 ml of sterile infant milk formula (Cow & Gate Premium 1), to achieve a final concentration of 20% (v/v), was aseptically added to the PCA and mixed thoroughly. The medium was then dispensed into Petri dishes and allowed to solidify and dry at room temperature (Alsonosi 2017; Cairns et al., 2025). All test isolates were inoculated onto XLD agar and PIF agar plates and incubated at 37°C for 24 hours. Qualitatively, capsule production was assessed based on colony morphology and the presence of hypermucoid phenotypes. Antimicrobial susceptibility testingIso-Sensitest agar (ISA) (Oxoid, UK) was prepared by dissolving 31.4 g of powder in 1 l of distilled water, followed by boiling to ensure complete dissolution. The medium was sterilised by autoclaving at 121°C for 15 minutes, cooled to approximately 50°C, and poured into Petri dishes. Antimicrobial susceptibility testing was performed using the Kirby and Bauer (disc diffusion) method in accordance with the British Society for Antimicrobial Chemotherapy (BSAC) guidelines (2015). The antibiotics tested represented five major classes: carbapenems (meropenem 10 µg and imipenem 10 µg), cephalosporins (Cefotaxime 30 µg, Ceftriaxone 30 µg, Ceftazidime 10 µg, and Ceftizoxime 30 µg), penicillins (ampicillin 10 µg), fluoroquinolones (ciprofloxacin 5 µg), and aminoglycosides (gentamicin 10 µg). Extended spectrum β-lactamase (ESBL) production was assessed using combination discs containing cefotaxime/clavulanic acid 30/10 µg and ceftazidime/clavulanic acid 30/10 µg, following the manufacturer’s instructions (Mast Diagnostics, Bootle, UK). Escherichia coli ATCC 10418 was included as a susceptible quality control isolate. For inoculum preparation, four colonies from a fresh TSA culture were suspended in 3 ml of sterile normal saline, and the turbidity was adjusted to match a 0.5 McFarland standard. The bacterial suspension was evenly spread onto ISA plates using a sterile swab, after which antibiotic discs were placed onto the agar surface. Plates were incubated at 37°C for 20 hours. Zones of inhibition were measured and interpreted in accordance with BSAC criteria 2015 (BSAC 2015). Whole genome sequencing and bioinformatic analysisGenomic DNA was extracted from all isolates using the GenElute™ Bacterial Genomic DNA Kit (Sigma Aldrich), following the manufacturer’s protocol. DNA libraries were prepared using the Nextera XT library preparation kit (Illumina, San Diego, CA), and whole genome sequencing was performed on an Illumina MiSeq platform using the MiSeq reagent kit v3, generating 250 bp paired-end reads. The reads were assembled de novo using Spades. Genome annotation was performed using Prokka (http://www.vicbioinformatics.com/software.prokka.shtml). Speciation of the isolates was confirmed by core genome alignment using parsnp, and the phylogenetic trees were visualised with Gingr, both of which belong to the Harvest package. The problematic threat factors genes of representative isolates were identified by using Virulence Finder 1.5 and the Artemis genome browser (Carver et al., 2005). In addition, Whole genome sequencing (WGS) was performed for the presented isolates to determine their O serotyping profile and to ascertain whether there is an association between O serotyping and K serotyping in K. pneumoniae infections. The genomes of isolates were visualised using the WebAct comparative tool and the Artemis comparative tool (Carver et al., 2005). Additional genome comparisons and sequence analyses were performed using BLAST searches against the NCBI GenBank database. Fully sequenced K. pneumoniae reference genomes NTUH K2044 and MGH 78578 were retrieved from GenBank and used for comparative analyses (https://www.ncbi.nlm.nih.gov/genbank/). Ethical approvalEthical approval from the NRES Committee East Midlands allowed the collection of feeding tubes, feces, and metadata but prohibited the storage of human tissue, including feces. ResultsA phenotypic assessment of capsular production was conducted for K. pneumoniae isolates using two different growth media: XLD agar and agar supplemented with PIF. As shown in Table 2, all isolates exhibited pronounced hypermucoid capsule formation on both media. Notably, isolates 1681 (ST111:01), 1699 (ST247:03), 1734 (ST526:02), 453 (ST105:04), 497 (ST147:012), and 1446 (ST37:02) produced substantially greater amounts of capsular material when grown on PIF agar compared with XLD agar. Table 2. Klebsiella pneumoniae serogroups and capsule production on XLD and PIF.
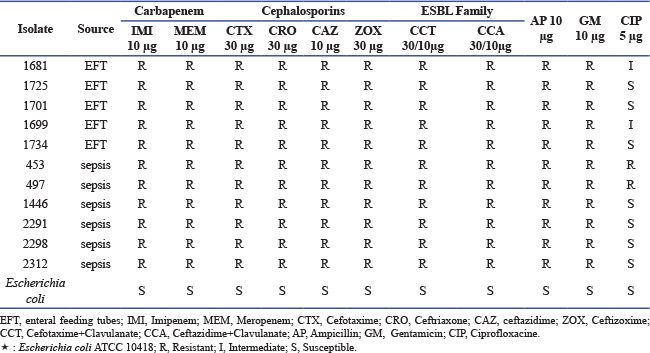
Antimicrobial susceptibility testing and ESBL activity assessment were performed for all isolates. The antimicrobial panel included representatives from five major antibiotic classes: carbapenems (meropenem and imipenem), cephalosporins (ceftazidime, cefotaxime, and cefpodoxime), fluoroquinolones (ciprofloxacin), penicillins (ampicillin), and aminoglycosides (gentamicin). ESBL production was evaluated using cefotaxime/clavulanic acid and ceftazidime/clavulanic acid combination assays. Escherichia coli NCTC 10418 was used as a quality control isolate and exhibited full susceptibility to all tested antibiotics. As presented in Table 3, all K. pneumoniae isolates demonstrated resistance to 10 of the 11 antibiotics tested and were susceptible only to ciprofloxacin (5 µg); however, isolates 453 and 497 also showed resistance to ciprofloxacin. All isolates were confirmed to be ESBL producers. Table 3. Antibiotic susceptibility of Klebsiella pneumoniae isolates from enteral feeding tubes and sepsis:
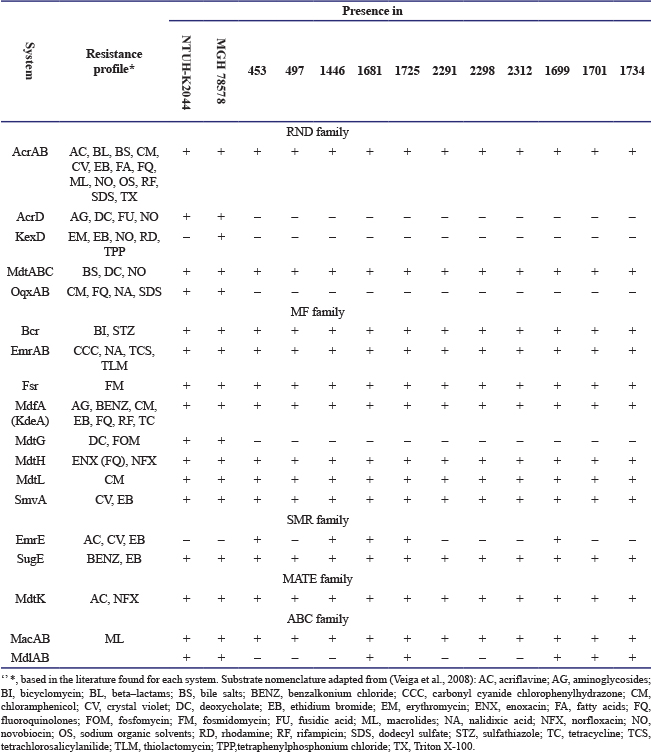
Whole genome sequencing WGS was performed to determine the sequence types and serogroups of the isolates. The analysis identified six distinct O-antigen serogroups, namely O1, O2, O3, O4, O8, and O12. Isolate 1681 (ST111) was assigned to serogroup O1, while three isolates (2312, 1446, and 1734) were classified as O2. Two isolates (1701 and 1699) belonged to serogroup O3. Isolate 453 was identified as O4, and isolate 2298 as O8. Serogroup O12 included two isolates (497 and 1725). The O-antigen serogroup of isolate 2291 could not be determined. With respect to capsular typing, serotype K1 was detected in isolates 1699 and 1701, whereas serotype K2 was identified in isolates 1681 and 1725. Serotype K5 was observed in isolates 453 and 497. Capsular serotypes were not detected in isolates 1446, 2291, 2298, 2312, and 1734 Table 2. Considerable variability was observed in the distribution of efflux pump genes among the isolates. Efflux pumps belonging to the RND family (AcrD, KexD, and OqxAB) and the major facilitator (MF) family (MdtG) were absent in all isolates. In contrast, the EmrE efflux pump, a member of the SMR family, was consistently detected in all isolates but was absent from the reference genomes K. pneumoniae NTUH K2044 and MGH 78578, Table 4. Table 4. Drug-related efflux pumps identified in compared Klebsiella pneumoniae.
DiscussionThe presence of K. pneumoniae in powdered infant formula or enteral feeding products represents a potential source of infection, particularly for immunocompromised individuals such as premature and hospitalised neonates. The risk of infection is heightened due to the increased susceptibility of these populations to nosocomial infections. Hospital-acquired infections remain a global challenge because of their clinical, financial, and ethical consequences, as well as the difficulty of effectively controlling outbreaks worldwide (World Health Organization 2011). Bacterial capsules are among the most important virulence factors and play a critical role in protecting bacteria from hostile environmental conditions, including phagocytosis, desiccation, acidic environments, and serum-mediated killing (Ogrodzki and Forsythe 2015; Holmes et al., 2021). In the present study, eleven K. pneumoniae isolates were examined to assess variations in capsular production on two different media (XLD and PIF agar). As shown in Table 2, substantial variability in capsule production was observed among the isolates, with hypermuco viscosity being more pronounced on PIF agar than on XLD agar. In particular, isolates 1681 (ST111:01), 1699 (ST247:03), 1734 (ST526:02), 453 (ST105:04), 497 (ST147:012), and 1446 (ST37:02) produced significantly greater amounts of capsular material on PIF agar. These findings indicate that media composition can influence capsular expression. Previous studies have shown that PIF components facilitate bacterial attachment to feeding tubes and promote bacterial growth and persistence (Acker et al., 2001). Additionally, bacteria grown in PIF have been reported to exhibit pronounced hypermuco viscosity, enabling growth on the media, a phenotype associated with increased virulence and potential neonatal health risks (Oyetibo et al., 2024). In the present study, six O-antigen serogroups were identified, namely O1, O2, O3, O4, O8, and O12 Table 2. Notably, isolates belonging to the O1 and O2 serotypes may pose a substantial risk to premature infants in neonatal intensive care units. These serotypes were detected across extended-spectrum β-lactamase-producing K. pneumonia and sepsis-associated isolates. Comparative analysis of virulence-associated traits among EFT and sepsis isolates indicated that isolate 1681 (O1: K2), isolated from an EFT case in Jordan, exhibited a higher virulence potential than the sepsis-associated isolates examined. Notably, a clear association was observed between sequence type ST247 and the O3 O-antigen and K1 capsular serotype (ST247:O3: K1). The introduction of antibiotics represents one of the most significant advances in medical history, substantially reducing morbidity and mortality associated with infectious diseases (UNDESA 2015). However, the rapid emergence and global spread of antimicrobial resistance pose a serious public health concern. In the present study, Table 3, most of K. pneumoniae isolates were resistant to 10 of the 11 antibiotics tested, with susceptibility observed only to ciprofloxacin, although two isolates (453 and 497) also exhibited ciprofloxacin resistance. Earlier studies reported higher susceptibility rates among K. pneumoniae isolates (Wang et al., 1998); however, these findings are now outdated. More recent investigations have documented widespread resistance to carbapenems, penicillins, and cephalosporins, with carbapenem resistance reported in more than 34% of K. pneumoniae isolates in a major New York medical centre (Kimani 2024). In 2017, the CDC identified carbapenem-resistant K. pneumoniae as one of the top three antimicrobial-resistant threats in the United States, accounting for approximately 7,900 infections and 520 deaths annually (Prevention 2017). Carbapenems are considered the treatment of choice for severe infections caused by ESBL-producing Enterobacteriaceae; however, the emergence of carbapenem-resistant isolates represents a critical therapeutic challenge due to limited alternative options (Pitout 2008; van Duin et al., 2014; Schweer et al., 2025). Although colistin has been proposed as an alternative treatment for carbapenem-resistant K. pneumoniae, increasing resistance and the plasmid-mediated spread of the mcr-1 gene are of growing concern. Consequently, colistin susceptibility testing is recommended for future investigations. All isolates examined in the current study were resistant to carbapenems, and the high level of multidrug resistance observed is alarming a potential risk to public health, despite partial susceptibility to ciprofloxacin. This resistance profile is likely associated with the high prevalence of β-lactamase (bla) genes and the presence of efflux pump systems. Notably, the EmrE efflux pump gene, belonging to the SMR family, was detected in isolates 453 (ST105:04), 1446 (ST37:02), 1681 (ST111:01), 1699 (ST247:03), and 1725 (ST111:012) but was absent in the reference isolates NTUH K2044 and MGH 78578, suggesting a potential role in enhanced antimicrobial resistance. Comparative genomic analysis indicates that the studied K. pneumoniae isolates harbour a broader repertoire of efflux pump genes and may exhibit increased multidrug resistance compared with the reference isolates. In this study, efflux pump-associated virulence factors were assessed solely on the basis of gene presence or absence; however, quantitative real time PCR would be necessary to confirm their expression and determine expression levels. Collectively, the findings of this study indicate that the K. pneumoniae isolates analysed harbour a concerning combination of high potential risk and elevated levels of antimicrobial resistance, representing a potential threat to public health. Importantly, heightened threats and antimicrobial resistance were not confined to isolates recovered from sepsis cases. Indeed, isolates 1681 and 1699 (ST11), as well as isolate 1734 (ST526), isolated from EFT cases, demonstrated greater virulence potential than isolates obtained from neonates with sepsis. ConclusionThe K. pneumoniae isolates examined in this study exhibited pronounced capsular production and extensive multidrug resistance, including universal carbapenem resistance and confirmed ESBL activity. Enhanced capsule expression on powdered infant formula agar implies that enteral feeding environments may serve as favorable niches for bacterial proliferation and persistence. Whole genome sequencing revealed substantial genetic diversity across sequence types and serogroups, with resistance and virulence determinants distributed across multiple lineages. The expanded efflux pump gene repertoire, particularly the presence of emrE, may contribute to the heightened resistance and persistence of these isolates. Collectively, these findings highlight a significant potential risk to neonatal health and underscore the need for strengthened infection control and ongoing genomic surveillance. AcknowledgmentThe authors thank the Libyan Biotechnology Research Centre (Tripoli, Libya) for their unlimited support of this study. Conflict of interestThe authors declare that there is no conflict of interest. FundingThe authors declare that this research received no external funding. Authors' contributionsM. Saad and K. Ibrahim conceived and designed the study. M. Saad, O. Elhenshir, A. Zaghdani, and B. Elmihub conducted the research. Data analysis was performed by M. Saad, M. Agena, A. Alsonosi, and N. Saiennaser. All authors contributed to revising the manuscript, reviewed and approved the final version, and accept full responsibility for the integrity and accuracy of the work. Data availabilityThe datasets used and/or analysed during the current study are available from the corresponding author upon request. References(n.d). WHO bacterial priority pathogens list 2024: Bacterial pathogens of public health importance to guide research, development and strategies to prevent and control antimicrobial resistance. Available via https://www.who.int/publications/i/item/9789240093461 (Accessed 9 December 2024). Alsonosi, A. 2017. Identification of physiological and virulence traits of clinical strains of Cronobacter malonaticus. Thesis. Available via https://irep.ntu.ac.uk/id/eprint/31949/1/Thesis%20Abdlrhman.pdf British Society for Antimicrobial Chemotherapy guidelines (BSAC). 2015. British Society for Antimicrobial Chemotherapy. Available via https://bsac.org.uk/susceptibility/ Davin-Regli, A., Masi, M., Bialek, S., Nicolas-Chanoine, M.H., and Pagès, J.M. 2016. Antimicrobial drug efflux pumps in Enterobacter and Klebsiella. Efflux-Mediated Antimicrobial Resistance in Bacteria: Mechanisms, Regulation and Clinical Implications. Fursova, N.K., Astashkin, E.I., Ershova, O.N., Aleksandrova, I.A. Savin, T.S., Novikova, et al. 2021. Multidrug-resistant Klebsiella pneumoniae causing severe infections in the Neuro-ICU Antibiotics 10(8), 1–17; doi: 10.3390/antibiotics10080979 Cairns, S., Gilpin, D. and Cameron, S.J.S. 2025. Infant formula milk shows microbiological contaminants that are not removed using recommended preparation methods. J. Appl. Microbiol. 136(11), lxaf267; doi:10.1093/jambio/lxaf267 Carver, T.J., Rutherford, K.M., Berriman, M., Rajandream, M.A., Barrell, B.G. and Parkhill, J. 2005. ACT: the Artemis comparison tool. Bioinformatics 21(16), 3422–3423; doi:10.1093/bioinformatics/bti553 Choby, J.E., Howard-Anderson, J. and Weiss, D.S. 2020. Hypervirulent Klebsiella pneumoniae - clinical and molecular perspectives. J. Intern. Med. 287(3), 283–300; doi:10.1111/joim.13007 Davin-Regli, A., Masi, M., Bialek, S., Nicolas-Chanoine, M. H. and Pagès, J. M. 2016. Antimicrobial drug efflux pumps in Enterobacter and Klebsiella. In Efflux-Mediated Antimicrobial Resistance in Bacteria: Mechanisms, Regulation and Clinical Implications. HYPERLINK "https://doi.org/10.1007/978-3-319-39658-3_11" doi: 10.1007/978-3-319-39658-3_11 De Oliveira, D.M.P., Forde, B.M., Kidd, T.J., Harris, P.N.A., Schembri, M.A., Beatson, S.A., Paterson, D.L. and Walker, M.J. 2020. Antimicrobial resistance in ESKAPE pathogens. Clin. Microbiol. Rev. 33, 181. Fang, C.T., Shih, Y.J., Cheong, C.M. and Yi, W.C. 2016. Rapid and Accurate Determination of Lipopolysaccharide O-Antigen Types in Klebsiella pneumoniae with a Novel PCR-Based O-Genotyping Method. J. Clin. Microbiol. 54(3), 666–675; doi:10.1128/JCM.02494-15 Follador, R., Heinz, E., Wyres, K.L., Ellington, M.J., Kowarik, M., Holt, K.E. and Thomson, N.R. 2016. The diversity of Klebsiella pneumoniae surface polysaccharides. Microbial. Genomics. 2(8), 73. doi: 10.1099/mgen.0.000073 Hansen, D.S., Skov, R., Benedí, J.V., Sperling, V. and Kolmos, H.J. 2002. Klebsiella typing: pulsed-field gel electrophoresis (PFGE) in comparison with O:K-serotyping. Clin. Microbiol. Infection. 8(7), 397–404; doi:10.1046/j.1469-0691.2002.00411.x Holmes, C.L., Anderson, M.T., Mobley, H.L.T. and Bachman, M.A. 2021. Pathogenesis of gram-negative bacteremia. Clin. Microbiol. Rev. 34, 234; doi:10.1128/cmr.00234-20 Hsieh, P.F., Lin, T.L., Yang, F.L., Wu, M.C., Pan, Y.J., Wu, S.H. and Wang, J.T. 2012. Lipopolysaccharide O1 antigen contributes to the virulence in Klebsiella pneumoniae causing pyogenic liver abscess. PLos One. 7(3), 0033155. doi:10.1371/journal.pone.0033155 Joshi, S., Chaudhary, A., Kumar, R., Kushwaha, R.S. and Biswas, D. 2025. Efflux Pumps In Multidrug Drug resistant- Klebsiella pneumoniae: discovery of Efflux Inhibitors. AfrJBioSc 7(4), 752–764; doi:10.48047/AFJBS.7.4.2025.752-764 Kimani, R.W. 2024. A review of Carbapenems Resistance in the Current World. J. Med. Biomed. Lab. Sci. Res. 4(1), 5. Navon-Venezia, S., Kondratyeva, K. and Carattoli, A. 2017. Klebsiella pneumoniae: a major worldwide source and shuttle for antibiotic resistance. FEMS. Microbiol. Rev. 41, 252–275; doi:10.1093/femsre/fux013 Ni, R.T., Onishi, M., Mizusawa, M., Kitagawa, R., Kishino, T., Matsubara, F., Tsuchiya, T., Kuroda, T. and Ogawa, W. 2020. The role of RND-type efflux pumps in multidrug resistant mutants of Klebsiella pneumoniae. Sci. Rep. 10, 10876; doi:10.1038/s41598-020-67820-x Ogrodzki, P. and Forsythe, S. 2015. Capsular profiling of the Cronobacter genus and the association of specific Cronobacter sakazakii and C. malonaticus capsule types with neonatal meningitis and necrotizing enterocolitis. BMC. Genomics 16(1), 758–772. Oyetibo, O.B., Shittu, O.B. and Omemu, A.M. 2024. Isolation of Pathogenic Cronobacter Species as Bacteriological Risks Indicator in Powdered Infant Formula Available to Deprived Infants in Lagos Metropolis, Nigeria. Curr. Microbiol. 81, 360; doi:10.1007/s00284-024-03850-2 Piddock, L.J.V. 2006a. Clinically relevant chromosomally encoded multidrug resistance efflux pumps in bacteria. Clin. Microbiol. Rev. 19, 382–402; doi:10.1128/cmr.19.2.382-402.2006 Piddock, L.J.V. 2006b. Multidrug-resistance efflux pumps - not just for resistance. Nat. Rev. Microbiol. 4, 629–636; doi:10.1038/nrmicro1464 Piddock, L.J.V. 2006c. Multidrug-resistance efflux pumps: not just for resistance. Nat. Rev. Microbiol. 4, 629–636. Pitout, J.D. and Laupland, K.B. 2008. Extended-spectrum beta-lactamase-producing Enterobacteriaceae: an emerging public-health concern. Lancet. Infect. Dis. 8, 159–166. Ramos, P.I.P., Picão, R.C., De Almeida, L.G.P., Lima, N.C.B., Girardello, R., Vivan, A.C.P., Xavier, D.E., Barcellos, F.G., Pelisson, M., Vespero, E.C., Médigue, C., De Vasconcelos, A.T.R., Gales, A.C. and Nicolás, M.F. 2014. Comparative analysis of the complete genome of KPC-2-producing Klebsiella pneumoniae Kp13 reveals remarkable genome plasticity and a wide repertoire of virulence and resistance mechanisms. BMC Genomics 15, 54; http://www.biomedcentral.com/1471-2164/15/54 Santajit, S. and Indrawattana, N. 2016. Mechanisms of antimicrobial resistance in ESKAPE pathogens. Biomed. Res. Int. 2016, 2475067; doi: 10.1155/2016/2475067. Schweer, J., Peters, F. and Ahmad-Nejad, P. 2025. Rapid detection of carbapenemases in multiresistant Gram-negative strains: evaluation of two tests. Microbiol. Spectr. 13, e02789–e02724; doi:10.1128/spectrum.02789-24 Struve, C., Roe, C.C., Stegger, M., Stahlhut, S.G., Hansen, D.S., Engelthaler, D.M. and Krogfelt, A. 2015. Mapping the Evolution of Hypervirulent. mBio 6(4), 1–12; doi:10.1128/mBio.00630-15 The Centers for Disease Control and Prevention. 2017. Learn about cronobacter infection. Available via https://www.cdc.gov/features/cronobacter UNDESA. The Millennium Development Goals Report 2015. New York: United Nations Secretariat, 2015. Available via http://www.un.org/en/development/desa/publications/mdg-report-2015.html Van Acker, J., De Smet, F., Muyldermans, G., Bougatef, A., Naessens, A. and Lauwers, S. 2001. Outbreak of Necrotizing Enterocolitis Associated with Enterobacter sakazakii in Powdered Milk Formula. J. Clin. Microbiol. 39(1), 7–12; doi:10.1128/JCM.39.1.293 Van Duin, D., Kaye, K., Neuner, E. and Ra, B. 2014. Carbapenem-resistant. Diagnostic. Microbiol. Infect. Dis. 75(2), 115–120; doi:10.1016/j.diagmicrobio.2012.11.009 Veiga, D.F., Vicente, F.F., Nicolás, M.F. and Vasconcelos, A.T.R. 2008. Predicting transcriptional regulatory interactions with artificial neural networks applied to E. coli multidrug resistance efflux pumps. BMC. Microbiol. 8, 101–114. Wang, J.H., Liu, Y.C., Lee, S.S.J., Yen, M.Y., Wang, Y.S.C.H., Wann, S.R. and Lin, H.H. 1998. Primary liver abscess due to Klebsiella pneumoniae in Taiwan. Clin. Infect. Dis. 26, 1434–1438. World Health Organization. 2011. Core components for infection prevention and control programs. Assessment tools for IPC programmes. Prevention of hospital-acquired infections. Geneve, Switzerland: World Health Organization. Wyres, K.L. and Holt, K.E. 2018. Klebsiella pneumoniae as a key trafficker of drug resistance genes from environmental to clinically important bacteria. Curr. Opinion Microbiol. 45, 131–139; doi:10.1016/j.mib.2018.04.004 | ||
| How to Cite this Article |
| Pubmed Style Saad MT, Sifennasr NE, Agena MB, Elhenshir OS, Zaghdani AA, Alsonosi AM, Elmihub BR, Ibrahim KM. Genomic and phenotypic characterization of multidrug-resistant Klebsiella pneumoniae in neonatal care settings. J Microbiol Infect Dis. 2026; 16(1): 28-36. doi:10.5455/JMID.2026.v16.i1.5 Web Style Saad MT, Sifennasr NE, Agena MB, Elhenshir OS, Zaghdani AA, Alsonosi AM, Elmihub BR, Ibrahim KM. Genomic and phenotypic characterization of multidrug-resistant Klebsiella pneumoniae in neonatal care settings. https://www.jmidonline.org/?mno=303956 [Access: February 28, 2026]. doi:10.5455/JMID.2026.v16.i1.5 AMA (American Medical Association) Style Saad MT, Sifennasr NE, Agena MB, Elhenshir OS, Zaghdani AA, Alsonosi AM, Elmihub BR, Ibrahim KM. Genomic and phenotypic characterization of multidrug-resistant Klebsiella pneumoniae in neonatal care settings. J Microbiol Infect Dis. 2026; 16(1): 28-36. doi:10.5455/JMID.2026.v16.i1.5 Vancouver/ICMJE Style Saad MT, Sifennasr NE, Agena MB, Elhenshir OS, Zaghdani AA, Alsonosi AM, Elmihub BR, Ibrahim KM. Genomic and phenotypic characterization of multidrug-resistant Klebsiella pneumoniae in neonatal care settings. J Microbiol Infect Dis. (2026), [cited February 28, 2026]; 16(1): 28-36. doi:10.5455/JMID.2026.v16.i1.5 Harvard Style Saad, M. T., Sifennasr, . N. E., Agena, . M. B., Elhenshir, . O. S., Zaghdani, . A. A., Alsonosi, . A. M., Elmihub, . B. R. & Ibrahim, . K. M. (2026) Genomic and phenotypic characterization of multidrug-resistant Klebsiella pneumoniae in neonatal care settings. J Microbiol Infect Dis, 16 (1), 28-36. doi:10.5455/JMID.2026.v16.i1.5 Turabian Style Saad, Mohamed T., Nadia E. Sifennasr, Mahmoud B. Agena, Omar S. Elhenshir, Ahmed A. Zaghdani, Abdlrhman M. Alsonosi, Budour R. Elmihub, and Khaled M. Ibrahim. 2026. Genomic and phenotypic characterization of multidrug-resistant Klebsiella pneumoniae in neonatal care settings. Journal of Microbiology and Infectious Diseases, 16 (1), 28-36. doi:10.5455/JMID.2026.v16.i1.5 Chicago Style Saad, Mohamed T., Nadia E. Sifennasr, Mahmoud B. Agena, Omar S. Elhenshir, Ahmed A. Zaghdani, Abdlrhman M. Alsonosi, Budour R. Elmihub, and Khaled M. Ibrahim. "Genomic and phenotypic characterization of multidrug-resistant Klebsiella pneumoniae in neonatal care settings." Journal of Microbiology and Infectious Diseases 16 (2026), 28-36. doi:10.5455/JMID.2026.v16.i1.5 MLA (The Modern Language Association) Style Saad, Mohamed T., Nadia E. Sifennasr, Mahmoud B. Agena, Omar S. Elhenshir, Ahmed A. Zaghdani, Abdlrhman M. Alsonosi, Budour R. Elmihub, and Khaled M. Ibrahim. "Genomic and phenotypic characterization of multidrug-resistant Klebsiella pneumoniae in neonatal care settings." Journal of Microbiology and Infectious Diseases 16.1 (2026), 28-36. Print. doi:10.5455/JMID.2026.v16.i1.5 APA (American Psychological Association) Style Saad, M. T., Sifennasr, . N. E., Agena, . M. B., Elhenshir, . O. S., Zaghdani, . A. A., Alsonosi, . A. M., Elmihub, . B. R. & Ibrahim, . K. M. (2026) Genomic and phenotypic characterization of multidrug-resistant Klebsiella pneumoniae in neonatal care settings. Journal of Microbiology and Infectious Diseases, 16 (1), 28-36. doi:10.5455/JMID.2026.v16.i1.5 |