| Research Article | ||
J. Microbiol. Infect. Dis., (2026), Vol. 16(1): 23–27 Research Article The role of bacterial colonization in acute exacerbations of childhood bronchial asthma in Tripoli, LibyaHamida S. El-Magrahi*Department of Medical Laboratory Sciences, Faculty of Medical Technology, University of Tripoli, Tripoli, Libya *Corresponding Author: Hamida S. El-Magrahi. Department of Medical Laboratory Sciences, Faculty of Medical Technology, University of Tripoli, Tripoli, Libya. Email: Hamidasadkali [at] gmail.com Submitted: 18/10/2025 Revised: 30/12/2025 Accepted: 11/01/2026 Published: 27/01/2026 © 2026 Open Veterinary Journal
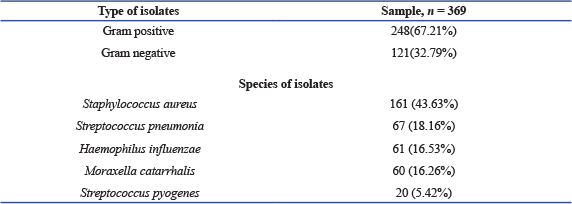
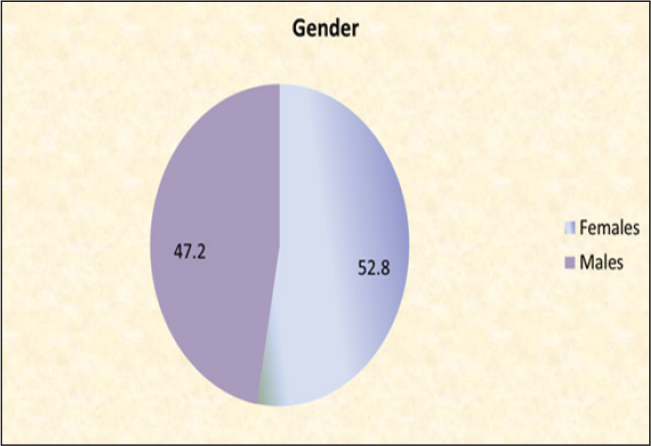
ABSTRACTBackground: Bronchial asthma is a complex, chronic condition of the airways and is one of the most prevalent pediatric illnesses worldwide and causing a significant, considerable amount of morbidity and mortality. Aim: This study aimed to identify bacterial species in the upper airway of asthmatic children, determine their antimicrobial susceptibility, and explore the role of bacterial infections in asthma exacerbations. Methods: This cross-sectional study included 369 children (195 females and 174 males) aged between 1 month and 15 years (mean age: 7.95 ± 3.95 years) from pediatric respiratory departments in hospitals across Tripoli, Libya, between January to April 2025. Demographic and clinical data were collected using structured questionnaires. Bacterial were samples collected nasopharyngeal swabs and sputum's. The bacterial isolates were identified using conventional identification techniques in accordance with normal microbiological protocols after the samples were cultivated. A Statistical analysis were conducted based on available data to examine microbial relationship to asthma in performed to examine associations between bacterial colonization, age, and gender. Results: Among 369 children, 195 (52.8%) were females, and 174 (47.2%) were males. The mean age of the children with asthma was (7.95 years ± 3.97 SD) (Range: month–15 years). Gram-positive bacteria accounted for 67.21% of isolates, while Gram-negative bacteria accounted for 32.79%. The most commonly isolated organism was Staphylococcus aureus (161; 43.63%), followed by Streptococcus pneumonia (67; 18.16% (Haemophilus influenza (61; 16.53%), Moraxella catarrhalis (60; 16.26%), and Streptococcus pyogenes (20: 5.42%). The highest culture positivity was observed in the 6–10 year age group, whereas the lowest was in infants aged 1–6 months. Most common organism grown in culture was S. aureus 161 (43.63%) followed by S. pneumonia 67 (18.16%), H. influenza 61 (16.53%), M. catarrhalis is 60 (16.26%) and S. pyogenes 20 (5.42%), Frequency distribution of isolated bacteria among children with asthma according to age, where the percentage of cultural positivity was highest in the age group 6–10 years, and the lowest rate of infection was recorded among age 1–6 months. Conclusion: Indicate these findings imply that bacterial colonization contributes to the pathogenesis of acute asthma and plays a significant role in Takes. Future research should focus on prevention and treatment strategies. Keywords: Asthma, Childhood, Bacterial colonization, Exacerbations in children, Antibiotic resistance. IntroductionAsthma is one of the most prevalent chronic illnesses in children. It is characterized by recurrent episodes of wheezing, dyspnea, chest tightness, and coughing, resulting from airway hyper responsiveness, associated with chronic inflammation (Toskala and Kennedy, 2015; Serebrisky and Wiznia, 2019). Globally, approximately 339 million people are affected by asthma worldwide (World Health Organisation, 2020). With a higher prevalence among children (9.5%) than adults (7.7%) (Loftus and Wise, 2016). Childhood asthma is in leading cause of morbidity and mortality (Asher and Pearce, 2014). The relationship between asthma and the microbial environment remain incompletely understood. Recent studies have increasingly focused on the role of microbes in the pathophysiology of asthma (Sullivan et al., 2016). Although microorganisms are known to influence airway inflammation, their specific contribution to asthma exacerbations is far less understood. Characterizing bacterial colonization in the airways of asthmatic patients could provide valuable insights into the mechanisms underlying airway inflammation and may pave the way for novel therapeutic approaches for chronic respiratory diseases such as asthma (Bokulich et al., 2016; Bush et al., 2017). Staphylococcus aureus has been frequently linked to both the onset and worsening of asthma (Langereis and de Jonge, 2015). A major Streptococcus pneumoniae infections are cause of mortality among children (Marangu and Zar, 2019). Heamophilus influenzae is among the one of the top five pathogenic bacteria that are most commonly found in young children (Eark et al., 2015). Additionally, H. influenzae can adhere to multiple receptors on the epithelium of the airways, which promotes increased inflammatory reactions (Huang and Boushey, 2014). Other respiratory pathogens, such as Mycoplasma pneumoniae and Chlamydia pneumoniae, have been detected in 5%–25% of children who experienced asthma flare-ups, while 20% of children who wheezed had Moraxella catarralis (Redinbo, 2014). Furthermore, the composition on the early-life microbiota's composition is significantly influenced by environmental factors such as formula feeding, birth mode, and antibiotic use (Jones et al., 2022). Although certain bacteria can cause acute respiratory infections by colonizing or infecting the airway, it remains unclear and hotly debated whether these specific bacterial infections are the actual cause of asthma flare-ups or serve as indicators of a higher risk of developing asthma (Simpson et al., 2016). Here, we discuss the available information about the potential contribution of infection to chronic asthma, with an emphasis on the potential role of bacteria. This study aimed to evaluate the upper airway microbial activity in children with asthma, to identify the type of bacteria and their profiles of susceptibility to the most widely used antimicrobial agents. Materials and MethodsStudy designThis study was conducted at the microbiology laboratory at the University Teaching Hospital, and pediatric respiratory departments in hospitals in Tripoli city, Libya, from January to April 2025. Study populationA total of 369 patients were included in the study, 195 females and 174 males, aged were one month and 15 years. The patients who participated in the study were from the respiratory department. Study procedureSample collection and transport- In this study, two samples were collected: The nasopharyngeal samples were obtained by inserting a swab approximately 1 inch into the nasopharyngeal and rotating it in a circular motion for 5 seconds each. Also, sputum samples were collected from the mouth in sterile plastic container after the patient took deep breath for 5 seconds. All samples were collected, early in the morning, immediately following the patient's awakening and prior to meals. Microorganism isolation and identificationSwab samples were inoculated onto chocolate agar and blood agar. After 24 hours of incubation at 37°C, the infected plates were checked for growth. If no growth, the plate was incubated for up to 48 hours before declaring a negative result (Bailey and Scott’s, 2022). Bacterial identification was performed using standard laboratory techniques. This is based on microscopic analysis, biochemical testing, hemolytic pattern, and colony morphology. Bacterial antibiotic susceptibility testingAccording to Clinical and Laboratory Standard Institute standards, antimicrobial susceptibility testing was carried out using the Kirby Bauer disc diffusion method (CLSI, 2020). Mueller Hinton agar plates were inoculated with pure cultures of the corresponding bacterial isolates. After 3–5 minutes of drying at room temperature, a set of antibiotic discs were placed on the plates. The antibiotic discs listed below, together with their corresponding concentrations, were used: Amikacin (AK, 30 µg); Tobramicin (TOB, 10 µg); Cefoxitin (CN, 30 µg); Cefiraxone (CRO, 30 µg); Imipenem (IPM, 10 µg); Meropenem (MEM, 10 µg); Eryhromycin (E, 10 µg); Sulphamethaxzol & Trimethoprim (SXT, 30 µg); Fosfomycin (FOX, 50 µg); Clindamycin (DA, 15 µg); Levofloxacin (LEV, 5 µg); Tigecyline (TIG, 15 µg); Oxacillin ( OX, 1 µg). Ethical approvalThe ethical research committee gave ethical approval for the study. The questionnaire was submitted to all patients to collect information such as age, gender, and health condition. ResultsOf the 369-study patient, 195 (52.8%) were females, and 174 (47.2%) were males as shown in Figure 1. The mean age of the children asthma was 7.95 years ± 3.97 SD (Range: 1 month–15 years). Out of the total isolates in the new paragraph Gram-positive and Gram-negative bacteria were identified in 248 cases (65.21%), while 121 (32.79%), respectively. Most common organism found in culture was S. aureus 161 (43.63%), followed by S. pneumonia 67 (18.16%) H. influenza 61 (16.53%), M. catarrhalis 60 (16.26%), and S. pyogenes 20 (5.42%), as shown in Table 1.
Fig. 1. Gender distribution of asthma cases among children. Table 1. Distribution of isolates microbial according to types and species.
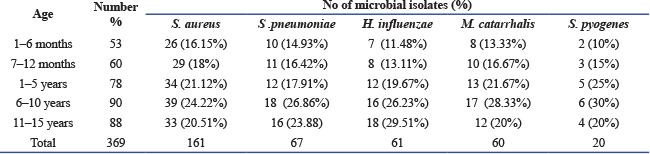
Bacterial isolates among children with asthma according to age is presented in Table 2. The highest rate of positive cultures was observed in the 6–10 year age group, while the lowest positivity rate of infection was recorded among infants aged 1–6 months. Table 2. Frequency distribution of isolated bacterial among asthma children according to age.
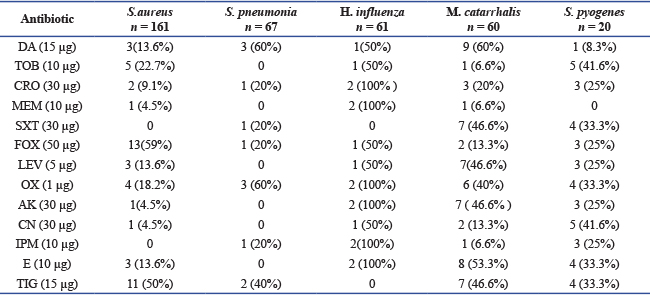
Antimicrobial Resistance Patterns from 65 asthmatic patients, showed significant bacterial growth after 24 hours, Gram-negative bacteria demonstrated high levels of antimicrobial resistance, as presented in Table 3. Thirteen antibiotics were tested for susceptibility. Pseudomonas aeruginosa exhibited 100% resistance all tested antimicrobials, Klebsiella pneumoniae showed 85% resistance to all antibiotics except (MEM), high resistance in gram positive, while S. aureus was 85% resistant to tested antimicrobial except (SXT, IPM) S. pyogenes was 85%, S. pneumonia showed moderate resistance 54% resistant to most antibiotics tested. Table 3. Prevalence and resistance of bacterial agents isolated from childhood asthma.
DiscussionAccording to the CDC, asthma is the most prevalent chronic illness in children. People with moderate to severe asthma are at risk for infection, which may trigger asthma attacks, pneumonia, and acute respiratory illnesses (Johnston, 2020). The frequency of childhood asthma varies significantly across countries (Serebrisky and Wiznia, 2019). Like everywhere else in the globe, asthma is a major health issue in Libya has revealed the presence of microorganisms in both the upper and lower respiratory tracts, which are linked to childhood asthma. Early childhood bacterial respiratory infections are strongly associated with the development of asthma. In our study, the frequency distribution of isolated bacteria according to age in Table 2. In healthy hosts, the bacteria often live as commensal organisms in the respiratory tract. However, it can cause illness in children younger than 2 years old, primarily because of the immune system's immaturity (Kumpitsch, et al., 2019). Studies from Turkey show that the frequency of asthma in children is between 8.6% and 12.6% (Jackson et al., 2020; Peters et al., 2020). Our study demonstrated the S. aureus was predominant in age groups. Age appears to have little effect on the association between S. aureus and wheezing diseases. These findings are consistent with investigation that the presence of Staphylococcus bacterium found in nasopharyngeal samples during the first 6 months of life is associated with a higher risk of childhood asthma and recurrent wheezing (Ali et al., 2021). Similary, neonatal colonization with H. influenza, S. pneumonia, and M. catarrhalis has been linked to an increased risk of recurrent wheeze and juvenile asthma (Tang et al., 2021). Collectively, these studies indicate that children's acute asthma episodes are linked to bacterial infections. which supports the findings of our study; however, the population of bacteria and the prevalence of them differs between the studies, this may be due to the environmental variations, study location, and differences in antibiotics resistance patterns. The identification of these bacteria in the airway of asthmatic. Future are warranted to studies clarify the rule of this bacteria and pathways in asthma pathogenesis. Future are warranted to studies clarify the rule of this bacteria and pathways in asthma pathogenesis. ConclusionA number of pathogens, including bacteria, are linked to the onset of asthma, and S. aureus is the primary cause of upper and lower respiratory infections, which are a common cause of acute sickness in children and an exacerbation of asthma. These pathogens are resistant to the most widely used antibiotics. AcknowledgmentI am grateful to the editors and anonymous reviewers for their thoughtful recommendations regarding this research. Conflict of interestThe author declares that there is no conflict of intetest. FundingNo particular grant from governmental or private funding was received for this study. Data availabilityAll data were provided in the manuscript. ReferencesAli, M.T., Abduljawad, N.H. and Al-Zawi, A.S. 2021. The prevalence of asthma AND allergic diseases in school age children in Albayda-Libya. Ejpmr 8(2), 194–196. Asher, I. and Pearce, N. 2014. Global burden of asthma among children. Int. J. Tuberculosis Lung Dis. 18, 1269–1278. Bailey and Scott’s. 2021. Diagnostic microbiology. 15th ed. American Society for Clinical Pathology, Elsevier. Inc. Bokulich, N.A., Chung, J., Battaglia, T., Henderson, N., Jay, M., Li, H., D. Lieber, A., Wu, F., Perez-Perez, G.I., Chen, Y., Schweizer, W., Zheng, X., Contreras, M., Dominguez-Bello, M.G. and Blaser, M.J. 2016. Antibiotics, birth mode, and diet shape microbiome maturation during early life. Sci. Transl. Med. 8, 343–382. Bush, A., Fleming, L. and Saglani, S. 2017. Severe asthma in children. Respirology 22, 886–897. CLSI, 2020. Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing. 30th ed. Wayne, PA, USA. Earl, C.S., An, S.Q. and Ryan, R.P. 2015. The changing face of asthma and its relation with microbes. Trends Microbiol. 23(7), 408–418. Huang, Y.J. and Boushey, H.A. 2014. The microbiome and asthma. Ann. Am. Thorac. Soc. 1, 48–51. Jackson, D.J., Busse, W.W., Bacharier, L.B., Kattan, M., O’Connor, G.T., Wood, R.A., Visness, C.M., Durham, S.R., Larson, D., Esnault, S., Ober, C., Gergen, P.J., Becker, P., Togias, A., Gern, J.E. and Altman, M.C. 2020. Association of respiratory allergy, asthma, and expression of the SARS-CoV-2 receptor ACE2. J. Allergy Clin. Immunol. 146(1), 203–206. Johnston, S.L. 2020. Asthma and COVID-19: is asthma a risk factor for severe outcomes?. Allergy 75(7), 1543–1545. Jones, H., Lawton, A. and Gupta, A. 2022. Asthma attacks in children challenges and opportunities. Indian J. Pediatrics 89(4), 373–377. Kumpitsch, C., Koskinen, K., Schöpf, V. and Moissl-Eichinger, C. 2019. The microbiome of the upper respiratory tract in health and disease. BMC. Biol. 17(87), 2–20. Langereis, J.D. and De Jonge, M.I. 2015. Invasive disease caused by non-typeable Haemophilus influenzae. Emerg. Infect. Dis. 21, 1711–1718. Loftus, P.A. and Wise, S.K. 2016. Epidemiology of asthma. Curr. Opinion Otolaryngol. & Head Neck Surg. 24(3), 245–249. Marangu, D. and Zar, H.J. 2019. Childhood pneumonia in low-and-middle-income countries: an update. Paediatr. Respir. Rev. 32, 3–9. Peters, M.C., Sajuthi, S., Deford, P., Christenson, S., Rios, C.L. and Montgomery, M.T. 2020. SARSCOV-2 related genes in sputum cells in asthma: relationship to demographic features and corticosteroids. Am. J. Respir. Crit. Care Med. 202(1), 83–90. Redinbo, M.R. 2014. The microbiota chemical symbiosis, and human disease. J. Mol. Biol. 426, 3877–3891. Serebrisky, D. and Wiznia, A. 2019. Pediatric asthma: a global epidemic. Ann. Global. Health. 85(1), 1–6. Simpson, J.L., Daly, J., Baines, K.J., Yang, I.A., Upham, J.W., Reynolds, P.N., Hodge, S., James, A.L., Hugenholtz, P., Willner, D. and Gibson, P.G. 2016. Airway dysbiosis: haemophilus influenzae and Tropheryma in poorly controlled asthma. Euro. Respir. J. 47, 792. Sullivan, A., Hunt, E., Macsharry, J. and Murphy, D.M. 2016. The microbiome and the pathophysiology of asthma. Respir. Res. 17(1), 163. Tang, H.H.F., Lang, A., Teo, S.M., Judd, L.M., Gangnon, R., Evans, M.D., Lee, K.E., Vrtis, R., Holt, P.G., Lemanske, R.F., Jackson, D.J., Holt, K.E., Inouye, M. and Gern, J.E. 2021. Developmental patterns in the nasopharyngeal microbiome during infancy are associated with asthma risk. J. Allergy Clin. Immunol. 147(5), 1683–1691. Toskala, E. and Kennedy, D. 2015. Asthma risk factors. Int. Forum. Allergy. Rhinol. 5, 11–16. World Health Organisation. Asthma. 2020. Available via https://www.who.int/news-room/fact-sheets/detail/asthma. | ||
| How to Cite this Article |
| Pubmed Style Hamida S. El-Magrahi. The role of bacterial colonization in acute exacerbations of childhood bronchial asthma in Tripoli, Libya. J Microbiol Infect Dis. 2026; 16(1): 23-27. doi:10.5455/JMID.2026.v16.i1.4 Web Style Hamida S. El-Magrahi. The role of bacterial colonization in acute exacerbations of childhood bronchial asthma in Tripoli, Libya. https://www.jmidonline.org/?mno=291258 [Access: January 29, 2026]. doi:10.5455/JMID.2026.v16.i1.4 AMA (American Medical Association) Style Hamida S. El-Magrahi. The role of bacterial colonization in acute exacerbations of childhood bronchial asthma in Tripoli, Libya. J Microbiol Infect Dis. 2026; 16(1): 23-27. doi:10.5455/JMID.2026.v16.i1.4 Vancouver/ICMJE Style Hamida S. El-Magrahi. The role of bacterial colonization in acute exacerbations of childhood bronchial asthma in Tripoli, Libya. J Microbiol Infect Dis. (2026), [cited January 29, 2026]; 16(1): 23-27. doi:10.5455/JMID.2026.v16.i1.4 Harvard Style Hamida S. El-Magrahi (2026) The role of bacterial colonization in acute exacerbations of childhood bronchial asthma in Tripoli, Libya. J Microbiol Infect Dis, 16 (1), 23-27. doi:10.5455/JMID.2026.v16.i1.4 Turabian Style Hamida S. El-Magrahi. 2026. The role of bacterial colonization in acute exacerbations of childhood bronchial asthma in Tripoli, Libya. Journal of Microbiology and Infectious Diseases, 16 (1), 23-27. doi:10.5455/JMID.2026.v16.i1.4 Chicago Style Hamida S. El-Magrahi. "The role of bacterial colonization in acute exacerbations of childhood bronchial asthma in Tripoli, Libya." Journal of Microbiology and Infectious Diseases 16 (2026), 23-27. doi:10.5455/JMID.2026.v16.i1.4 MLA (The Modern Language Association) Style Hamida S. El-Magrahi. "The role of bacterial colonization in acute exacerbations of childhood bronchial asthma in Tripoli, Libya." Journal of Microbiology and Infectious Diseases 16.1 (2026), 23-27. Print. doi:10.5455/JMID.2026.v16.i1.4 APA (American Psychological Association) Style Hamida S. El-Magrahi (2026) The role of bacterial colonization in acute exacerbations of childhood bronchial asthma in Tripoli, Libya. Journal of Microbiology and Infectious Diseases, 16 (1), 23-27. doi:10.5455/JMID.2026.v16.i1.4 |